On-demand access (discovery) for widely distributed web-based data, publication and software resources, with highly refined search tools. There are two broad classes of product we envision:
- Tools to help researchers Publish their data, making it discoverable
- Tools to help researchers Discover published data.
These tools are designed to facilitate data sharing, use and discovery. ReproNim is developing an integrated application, NeuroBLAST, that will powerfully enable user-specified search and publish functions to data repositories, published studies, versioned software packages, study-related questions, and content.
Read more.
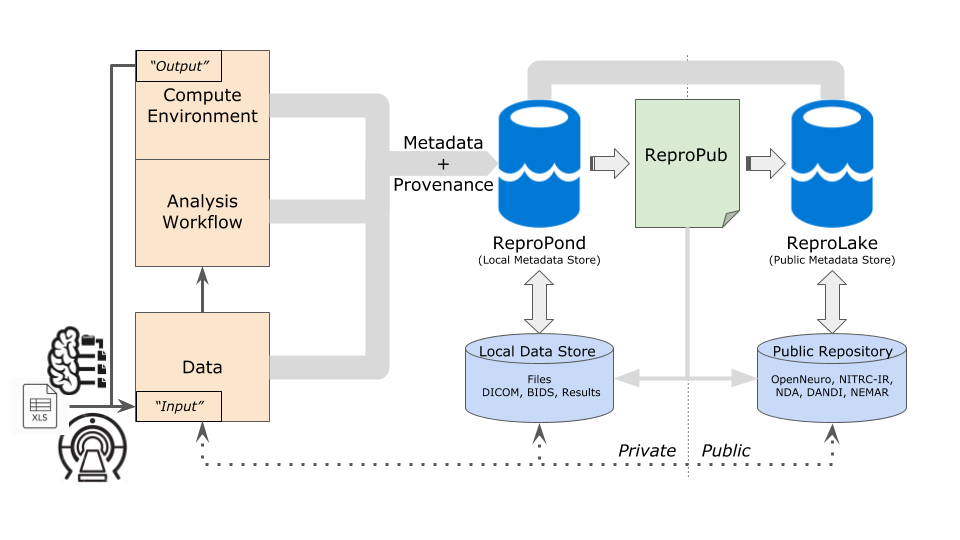
Reproducible description based on recording, reporting, and reusing experimenter procedures from start to finish. Machine-captured experimental metadata from scanner (data acquisition) to methods, analyses, and results.
- Tools to help researchers record (describe) experimental procedures
- Tools to help researchers define and semantically describe workflow.
ReproNim is developing BrainVerse to help researchers manage, track and share information in a comprehensive format.
Read more.
Efficient management of data and scripts while capturing the
history and details of changes and analyses execution in a fashion
that facilitates inspection, comparison, modification, sharing, and
re-execution. We provide
- Support, maintenance, and promotion of platforms
(e.g., NeuroDebian) and
tools (e.g., Singularity
and DataLad) for efficient and
reproducible computation and data management
- Tools and materials to establish reproducible computation across
all stages of the neuroimaging research
ReproNim is developing ReproIn framework for turnkey
collection of BIDS datasets straight from the
scanner, and ReproMan to help researchers
track and manage computation resources that they have available and to
use them in a reproducible and scalable way.
Read more.